IN GENERAL,
we study biomolecules and their interactions by biophysical and bioinformatic tools. We focus on interactions driving specific recognition between biomolecules with potential diagnostic, medical or biotechnological use.
At the molecular level, the essence of the biological processes consists in interactions between biomolecules. Our goal is to understand mechanisms of their mutual recognition proteins and nucleic acid specific. We concentrate on determination of molecular structure and dynamics, study specific protein-protein and protein-DNA interactions. We take advantage of experimental and computational methods of protein engineering, structural biology and bioinformatics, and molecular modeling.
You can see the short video about our group
the Czech language video
PROJECTS RUNNING IN THE LABORATORY
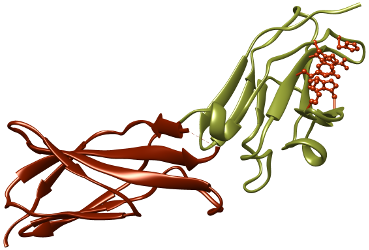
Cytokines
We study structure-function relationships of medically important signaling proteins crucial in immune response, cytokines. We study their structures and interactions and also try to modulate their function by developing de novo small protein binders.
Cytokines are small but important cell signaling proteins that are responsible for proper immune responses during inflammation, infection, trauma or cancer. Any error in their regulation causes serious autoimmune and/or allergic health disorders. We concentrate on the structural aspects of interactions between cytokines and extracellular parts of their corresponding receptors. In the last few years, we have worked on interferon-gamma and its two receptors, published several papers in this area and solved several crystal structures. Currently, we concentrate on medically important human proteins of the interleukin 10 family, the least known members of the family, IL-24, IL-26, IL-28, and IL-29.
We determine crystal structures and analyze binding characteristics of interactions between the cytokines and receptors. We elucidate the specificity of these interactions and modulate them by protein engineering techniques including directed in vitro evolution of a newly designed protein scaffold. We use ribosome and yeast display to de novo development of specific protein binders to FIL-10 proteins. We have developed a new small protein scaffold that has been trained to bind to interleukin 10.
The short video of the crystal structure of the extracellular part of receptor 2 of human interferon gamma (PDB ID: 5EH1).
These studies have been supported by the Czech Science Foundation projects P305-10-2184 (2010-14), 16-20507S (2016-2018), and 19-17398S (2019-2021).
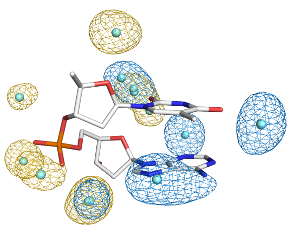
Bioinformatics
Our lab concentrates on structural analysis of nucleic acids, DNA and RNA, and of solvation around proteins and nucleic acids. Our bioinformatic research is based on original approaches and we develop our own analytical tools. The results are available on the website structbio.org.
Dinucleotide structures
Nucleic acids are structurally plastic molecules, and their biological functions are enabled by adaptation to their binding partners. We study structural aspects of their recognition by other molecules. Our work has contributed to understanding the structural behavior of their building blocks, dinucleotides. To identify structural polymorphisms of nucleic acids we analyzed hundreds of thousands of dinucleotide structures in hundreds of crystal structures. The work is summarized in several publications and the results are available on the website DNATCO.
Solvation
Part of the bioinformatic studies focuses on the structure of the solvation shell around biomolecules. First hydration shell of amino acids residues in proteins and DNA dinucleotides are studied from crystal structures. The results show how hydration depends on the amino acid or dinucleotide structure and sequence. The results have been published and are available on the webite WatAA.
Bioinformatic projects are supported by MEYS project ELIXIR-CZ: capacity building (CZ.02.1.01/0.0/0.0/16_013/0001777).
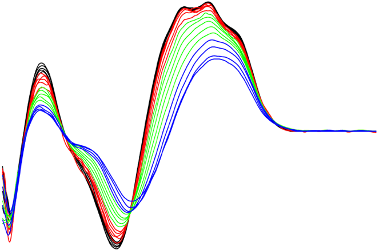
Single stranded DNA
Single stranded DNA (ssDNA) are known to play important yet largely unknown functions in the genome regulation. We study one specific class of bacterial ssDNA, so called Repetitive extragenic palindromes (REP) and their associated enzyme, tyrosine transposase (RAYT).
Our data reveal that many of the REP sequences may adopt multiple conformations. For several REP oligonucleotides studied experimentally, we obtained data consistent with formation of guanine tetraplexes as an alternative to the expected hairpin structures. It suggests that RAYTs confront a landscape of potential structural substates in dynamic equilibrium that could be selected, enriched, and/or induced via differential binding.
The project has been supported by the Czech Science Foundation, project P305-12-1801 (2012-2014), and by MEYS, project CIISB4HEALTH (CZ.02.1.01/0.0/16_013/0001776, 2018-2021).
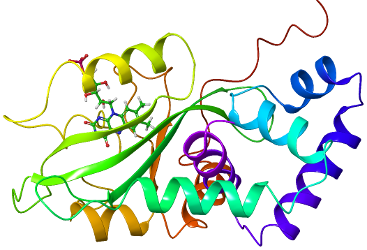
Dynamics of Biomolecules - ELIBIO Project
in collaboration with the ELI Beamlines research center
We explore new frontiers in light and optics to create breakthrough science about biomolecular dynamics at very high time resolution.
In collaboration with colleagues at the ELI-Beamlines infrastructure center in Dolní Břežany, we work on establishing an Interdisciplinary Centre of Excellence to exploit the photon beams of ELI-Beamlines in life sciences. An essential goal of the project is to understand photon-material interactions in extremely intense X-ray fields where new physics can be expected. We will use the new knowledge in studies on structure, function and dynamics in cells, organelles, and biomolecules to perform experiments that were impossible so far.
Supported by MEYS in project ELIBIO, Structural bioinformatics of biomolecular systems, (CZ.02.1.01/0.0/0.0/15_003/0000447, 2017-2022).

COOPERATION
The lab is a part of IBT, located at the BIOCEV center, and actively participates at ELIXIR and Instruct-ERIC infrastructural projects.
The Laboratory of Biomolecular Recognition is an intergral part of the Institute of Biotechnology of the Czech Academy of Sciences (IBT). The whole IBT is located at the center BIOCEV in Vestec. The newly built center is well equipped to perform research in structural biology, biochemistry, molecular and cellular biology, and research using animal models.
In BIOCEV, the lab is a part of the Research program "Structural Biology and Protein Engineering”.
IBT operates an infrastructure Center of Molecular Structure, CMS. CMS has excellent equipment for biophysical characterization of biomolecules, for crystallization, diffraction and scattering experiments as well as for sophisticated experiments using mass spectrometry including analysis of chemical cross linking and H/D exchange.
CMS is a part of the Czech Instruct-ERIC infrastructure CIISB.
The lab actively participates in the European infrastructure ELIXIR, especially in the area of structural bioinformatics where BS is a member of the Executive Committee of the 3D-Bioinfo community.
